Of course! Here is a comprehensive guide to using GaussianNB in Python, including a detailed explanation, code examples, and best practices.
What is Gaussian Naive Bayes?
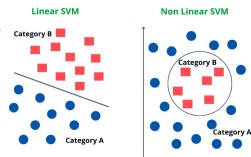
GaussianNB is a classification algorithm based on Bayes' Theorem with a "naive" assumption of independence between every pair of features.
Key Concepts:
-
Bayes' Theorem: It calculates the probability of a label given some features. The formula is: $P(y | x_1, x_2, ..., xn) = \frac{P(y) \prod{i=1}^{n} P(x_i | y)}{P(x_1, x_2, ..., x_n)}$
- $P(y | x_1, ..., x_n)$: Posterior probability (what we want to find: the probability of a class
ygiven the features). - $P(y)$: Prior probability (the probability of a class
yoccurring, regardless of features). - $P(x_i | y)$: Likelihood (the probability of a feature
x_ioccurring, given that the data point belongs to classy). - $P(x_1, ..., x_n)$: Evidence (the probability of the features occurring, which is constant for all classes and can be ignored during comparison).
- $P(y | x_1, ..., x_n)$: Posterior probability (what we want to find: the probability of a class
-
"Naive" Assumption: The algorithm assumes that all features are independent of each other given the class label. In reality, features are often correlated. This simplification makes the calculation much easier and works surprisingly well in many real-world scenarios.
 (图片来源网络,侵删)
(图片来源网络,侵删) -
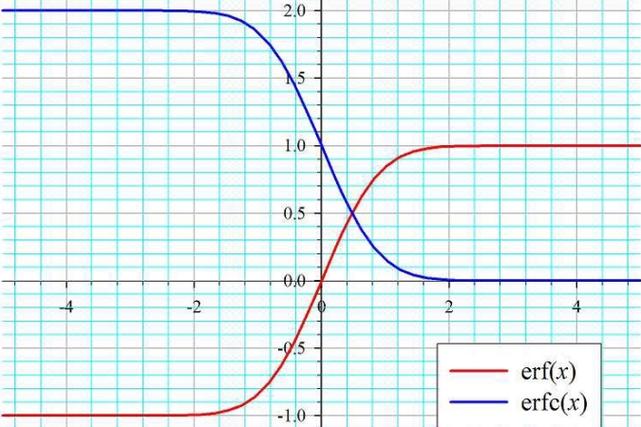
"Gaussian" Part: This specific version of Naive Bayes assumes that the values of each feature for a given class are drawn from a Gaussian (Normal) distribution. This means it works best with continuous numerical data. For each feature and each class, the algorithm calculates:
- The mean ($\mu$)
- The variance ($\sigma^2$)
When predicting the class for a new data point, it calculates the probability of that data point belonging to each class using the Gaussian probability density function for each feature, multiplies them together (due to the independence assumption), and picks the class with the highest probability.
When to Use GaussianNB?
It's an excellent choice when:
- You have a medium to large-sized dataset.
- Your features are continuous numerical data.
- You need a fast, simple, and interpretable model.
- The "naive" independence assumption is reasonably valid for your problem.
Implementation with scikit-learn
scikit-learn provides a simple and efficient implementation of GaussianNB in the naive_bayes module.
Step 1: Import Necessary Libraries
import numpy as np import matplotlib.pyplot as plt import seaborn as sns from sklearn.model_selection import train_test_split from sklearn.datasets import make_classification from sklearn.naive_bayes import GaussianNB from sklearn.metrics import accuracy_score, confusion_matrix, classification_report
Step 2: Create or Load Data
Let's create a synthetic dataset for a clear example. We'll use make_classification to generate a dataset with 2 classes and several informative features.
# Generate a synthetic dataset
# n_samples: number of data points
# n_features: number of features
# n_informative: number of features that are actually useful for classification
# n_redundant: number of features that are linear combinations of informative features
# n_classes: number of classes
# n_clusters_per_class: number of clusters for each class
X, y = make_classification(
n_samples=1000,
n_features=10,
n_informative=5,
n_redundant=2,
n_classes=2,
random_state=42
)
print("Shape of features (X):", X.shape)
print("Shape of labels (y):", y.shape)
print("\nFirst 5 rows of features:\n", X[:5])
print("\nFirst 5 labels:", y[:5])
Step 3: Split Data into Training and Testing Sets
This is a crucial step to evaluate the model's performance on unseen data.
# Split the data into training and testing sets
# test_size=0.2 means 20% of the data will be used for testing
# random_state ensures that the split is the same every time we run the code
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.2, random_state=42
)
print(f"\nTraining set size: {X_train.shape[0]} samples")
print(f"Testing set size: {X_test.shape[0]} samples")
Step 4: Initialize and Train the GaussianNB Model
This is where the magic happens. The fit() method calculates the mean and variance of each feature for each class from the training data.
# Initialize the Gaussian Naive Bayes model
gnb = GaussianNB()
# Train the model on the training data
gnb.fit(X_train, y_train)
print("\nModel training complete.")
Step 5: Make Predictions on the Test Set
Now, we use the trained model to predict the class labels for the test data.
# Make predictions on the test data
y_pred = gnb.predict(X_test)
print("\nFirst 10 predictions:", y_pred[:10])
print("First 10 actual labels:", y_test[:10])
Step 6: Evaluate the Model's Performance
How well did our model do? Let's calculate some common metrics.
# Calculate the accuracy of the model
accuracy = accuracy_score(y_test, y_pred)
print(f"\nModel Accuracy: {accuracy:.4f}")
# Display the confusion matrix
# A confusion matrix shows the number of correct and incorrect predictions for each class.
cm = confusion_matrix(y_test, y_pred)
print("\nConfusion Matrix:")
print(cm)
# For a more visual representation of the confusion matrix
plt.figure(figsize=(6, 4))
sns.heatmap(cm, annot=True, fmt='d', cmap='Blues', xticklabels=['Class 0', 'Class 1'], yticklabels=['Class 0', 'Class 1'])
plt.xlabel('Predicted Label')
plt.ylabel('True Label')'Confusion Matrix')
plt.show()
# Display a detailed classification report
# Report includes precision, recall, and F1-score for each class.
report = classification_report(y_test, y_pred)
print("\nClassification Report:")
print(report)
Step 7: Understand the Model's Learned Parameters
One of the great things about GaussianNB is its interpretability. You can inspect the parameters it learned during training.
# The model calculates the mean and variance of each feature for each class.
# Shape: (n_classes, n_features)
print("\nClass Priors (P(y)): The probability of each class in the training set.")
print(gnb.class_prior_)
print("\nFeature Means (mu) for each class:")
print(gnb.theta_) # theta_ is the attribute for means
print("\nFeature Variances (sigma^2) for each class:")
print(gnb.var_) # var_ is the attribute for variances
Complete Runnable Code
Here is the full script from start to finish for easy copy-pasting.
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.model_selection import train_test_split
from sklearn.datasets import make_classification
from sklearn.naive_bayes import GaussianNB
from sklearn.metrics import accuracy_score, confusion_matrix, classification_report
# 1. Create a synthetic dataset
X, y = make_classification(
n_samples=1000,
n_features=10,
n_informative=5,
n_redundant=2,
n_classes=2,
random_state=42
)
# 2. Split the data
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
# 3. Initialize and train the model
gnb = GaussianNB()
gnb.fit(X_train, y_train)
print("Model training complete.")
# 4. Make predictions
y_pred = gnb.predict(X_test)
# 5. Evaluate the model
accuracy = accuracy_score(y_test, y_pred)
print(f"\nModel Accuracy: {accuracy:.4f}")
# Confusion Matrix
cm = confusion_matrix(y_test, y_pred)
print("\nConfusion Matrix:")
print(cm)
# Visualize Confusion Matrix
plt.figure(figsize=(6, 4))
sns.heatmap(cm, annot=True, fmt='d', cmap='Blues', xticklabels=['Class 0', 'Class 1'], yticklabels=['Class 0', 'Class 1'])
plt.xlabel('Predicted Label')
plt.ylabel('True Label')'Confusion Matrix')
plt.show()
# Classification Report
report = classification_report(y_test, y_pred)
print("\nClassification Report:")
print(report)
# 6. Inspect learned parameters
print("\n--- Learned Parameters ---")
print("Class Priors (P(y)):", gnb.class_prior_)
print("\nFeature Means (theta_):\n", gnb.theta_)
print("\nFeature Variances (var_):\n", gnb.var_)
Other Types of Naive Bayes in scikit-learn
GaussianNB is just one flavor. The choice depends on your data type:
| Model | When to Use | Assumption |
|---|---|---|
GaussianNB |
Continuous numerical data. | Features follow a normal distribution. |
MultinomialNB |
Discrete counts (e.g., word counts in text). | Features represent counts or frequencies. |
BernoulliNB |
Binary/Boolean features (0s and 1s). | Features are binary (e.g., presence/absence of a word). |
ComplementNB |
Imbalanced datasets. A variation of MultinomialNB that is often more effective. |
Features are counts. It complements the other class's probabilities. |